



What is DNA Fingerprinting? Definition, Important Terms, Process, Forensic Science Uses, and Limitations
DNA Fingerprinting is a modern genetic identification technique used to distinguish one individual from another by studying specific variable regions in DNA. It is one of the most important applications of molecular biology and biotechnology, widely used in forensic science, paternity testing, criminal investigation, disaster victim identification, and population genetics.
Since every person, except identical twins, has a unique DNA pattern, DNA fingerprinting has become a highly reliable scientific method for personal identification. The technique is based on the fact that although human DNA is almost identical across individuals, certain regions show variations called polymorphisms, which create a unique DNA profile for each person.
DNA Fingerprinting
DNA fingerprinting, also called DNA profiling, is a technique used to identify an individual by analysing unique patterns in their DNA. It compares selected regions of DNA that vary from person to person rather than sequencing the entire genome.
This method became a landmark development in biology and forensic science after it was introduced by Sir Alec Jeffreys in 1984. In India, major contributions to the development and use of DNA fingerprinting were made by Dr V. K. Kashyap and Dr Lalji Singh.
DNA fingerprinting is used in many real-life situations, such as:
identifying criminals from biological evidence
resolving paternity and family disputes
solving immigration-related identity issues
identifying dead bodies in accidents or disasters
studying genetic diversity among populations
Why is DNA Fingerprinting Needed?
Human beings share nearly 99.9% of their DNA, but around 0.1% of DNA differs among individuals. These differences are enough to make each person genetically unique.
If scientists had to compare the entire DNA sequence of two people every time, it would be extremely difficult, time-consuming, and expensive. DNA fingerprinting solves this problem by focusing only on specific highly variable repetitive DNA regions, making identification much faster and more practical.
Principle of DNA Fingerprinting
The principle of DNA fingerprinting is based on the presence of DNA polymorphisms in certain non-coding repetitive regions.
Key idea
Although most of the DNA sequence in humans is similar, some DNA regions differ in:
sequence
number of repeats
fragment length
These variable regions are inherited from parents and remain the same in all tissues of the same person. Because of this, DNA obtained from blood, saliva, bone, skin, sperm, hair follicles, and other tissues can be used for identification.
Why Does it Work?
DNA fingerprinting works because:
Each individual has a unique pattern of repetitive DNA
These polymorphisms are inherited
The same DNA pattern is found in all cells of a person
The probability of two unrelated people having the same DNA fingerprint is extremely low
DNA Polymorphism
DNA polymorphism refers to variation in the DNA sequence between individuals. These differences may arise from changes in the number of repeated DNA units or in specific nucleotide sequences.
DNA polymorphism forms the basis of:
DNA fingerprinting
genetic mapping
population studies
paternity testing
forensic identification
Because polymorphisms are inherited, they help trace biological relationships between individuals.
Repetitive DNA
DNA fingerprinting primarily examines specific DNA regions called repetitive DNA.
These are regions where a short DNA sequence is repeated many times. Repetitive DNA does not usually code for proteins, but it constitutes a significant portion of the human genome and exhibits high variation among individuals.
Important Features of Repetitive DNA
Feature | Description |
Nature | Repeated DNA sequences |
Function | Usually non-coding |
Occurrence | Present in large portions of the genome |
Importance | Shows high polymorphism |
Use | Basis of DNA fingerprinting |
These repetitive sequences can be separated from the bulk genomic DNA and studied for variation. Since the amount and arrangement of repeats differ from one individual to another, they produce distinct DNA patterns.
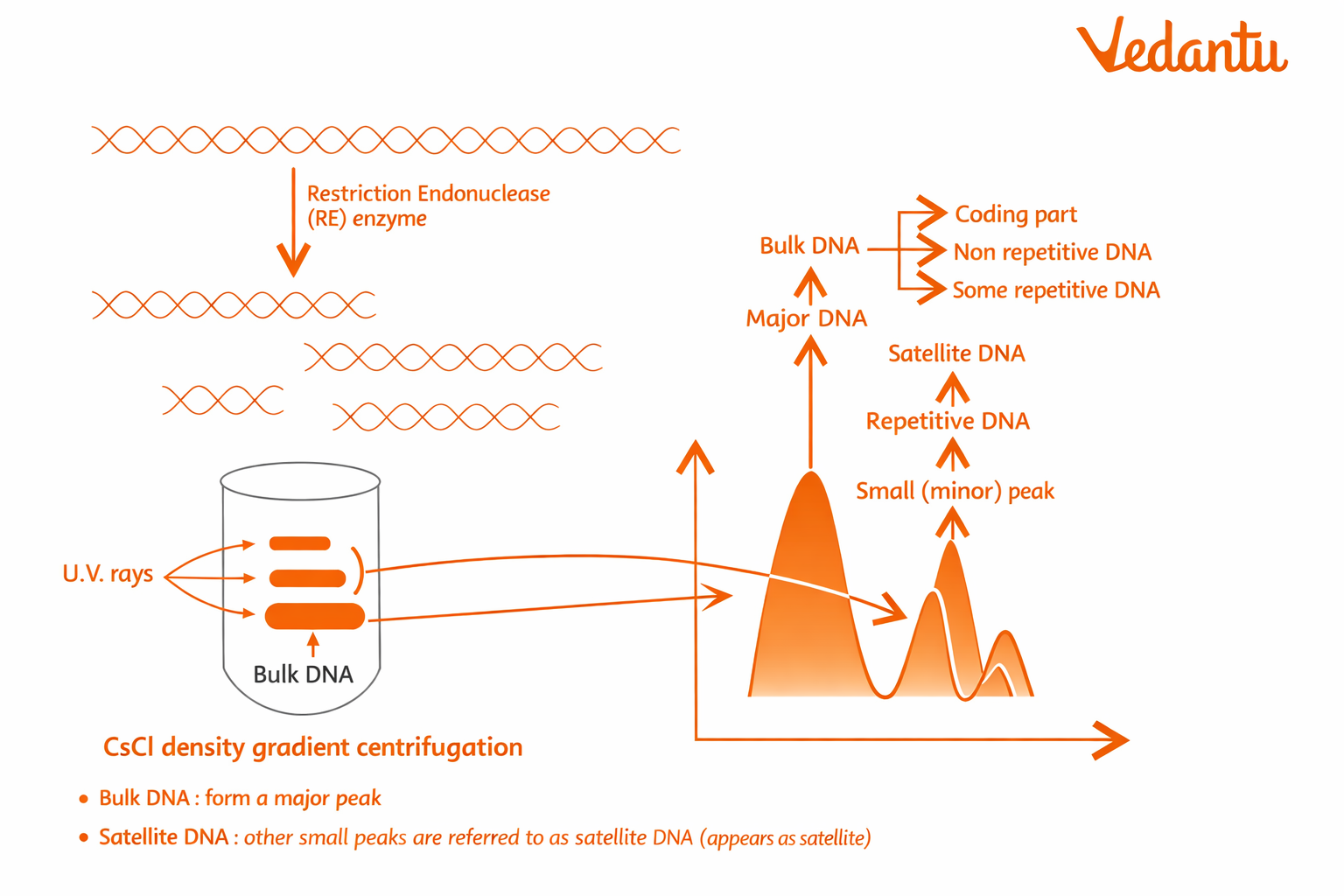
Why is Repetitive DNA Important in DNA Fingerprinting?
Repetitive DNA is highly useful because:
It varies greatly in a population
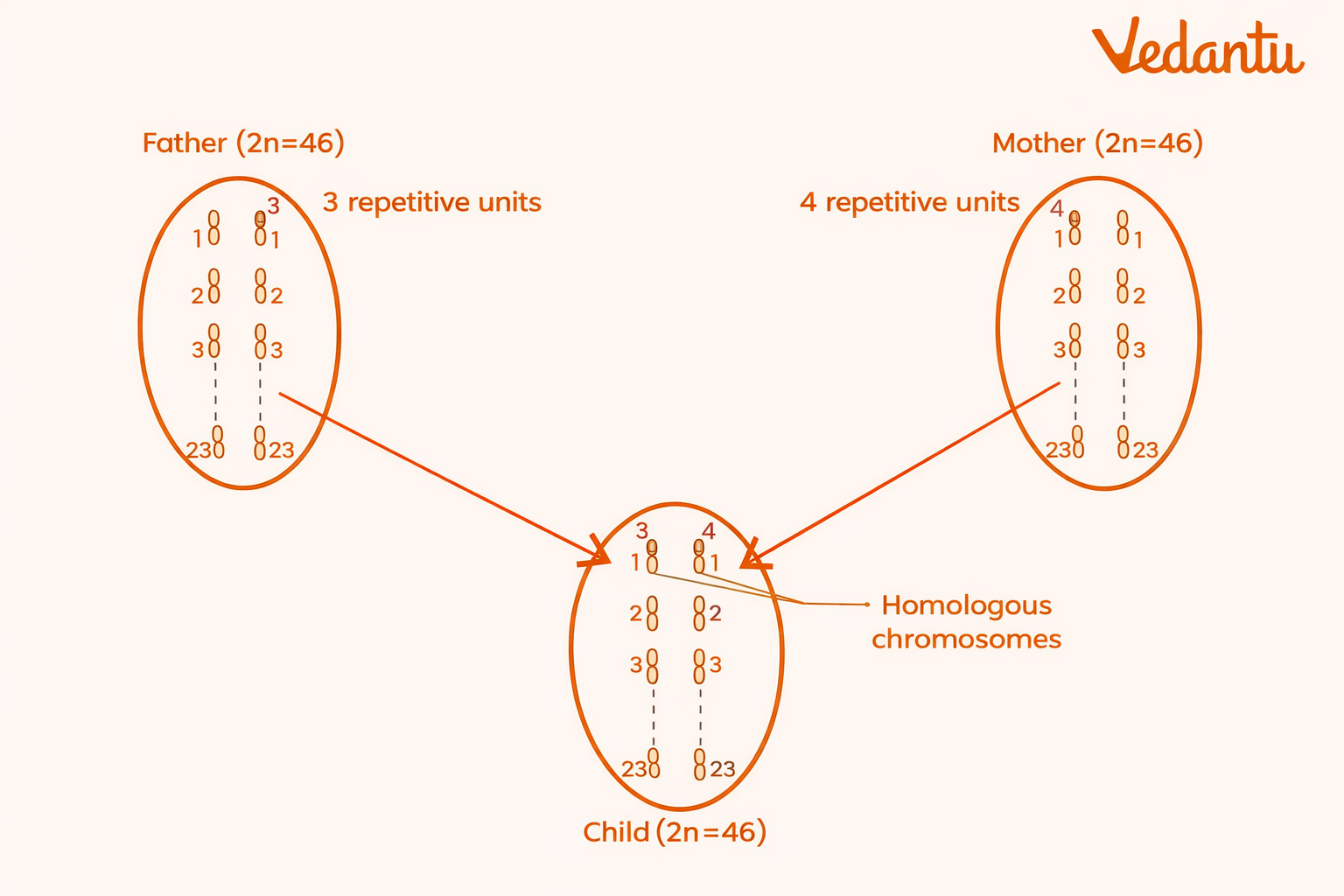
It can differ even between homologous chromosomes of the same individual
It remains the same in all tissues of one person
It is inherited from parents to children
This makes it ideal for forensic identification and paternity analysis. For example, DNA taken from blood at a crime scene can be matched with DNA from a suspect because both will show the same pattern if they come from the same person.
Satellite DNA
A large part of repetitive DNA belongs to a class called satellite DNA.
Satellite DNA is classified based on:
base composition
length of repeating unit
number of repetitive units
Main Categories of Satellite DNA
Type | Repeat Length | Other Name |
Microsatellites | 1 to 6 base pairs | SSR (Simple Sequence Repeats) |
Minisatellites | 11 to 60 base pairs | VNTR (Variable Number of Tandem Repeats) |
Satellite DNA is especially important because these regions are highly polymorphic and differ among individuals.
Microsatellites
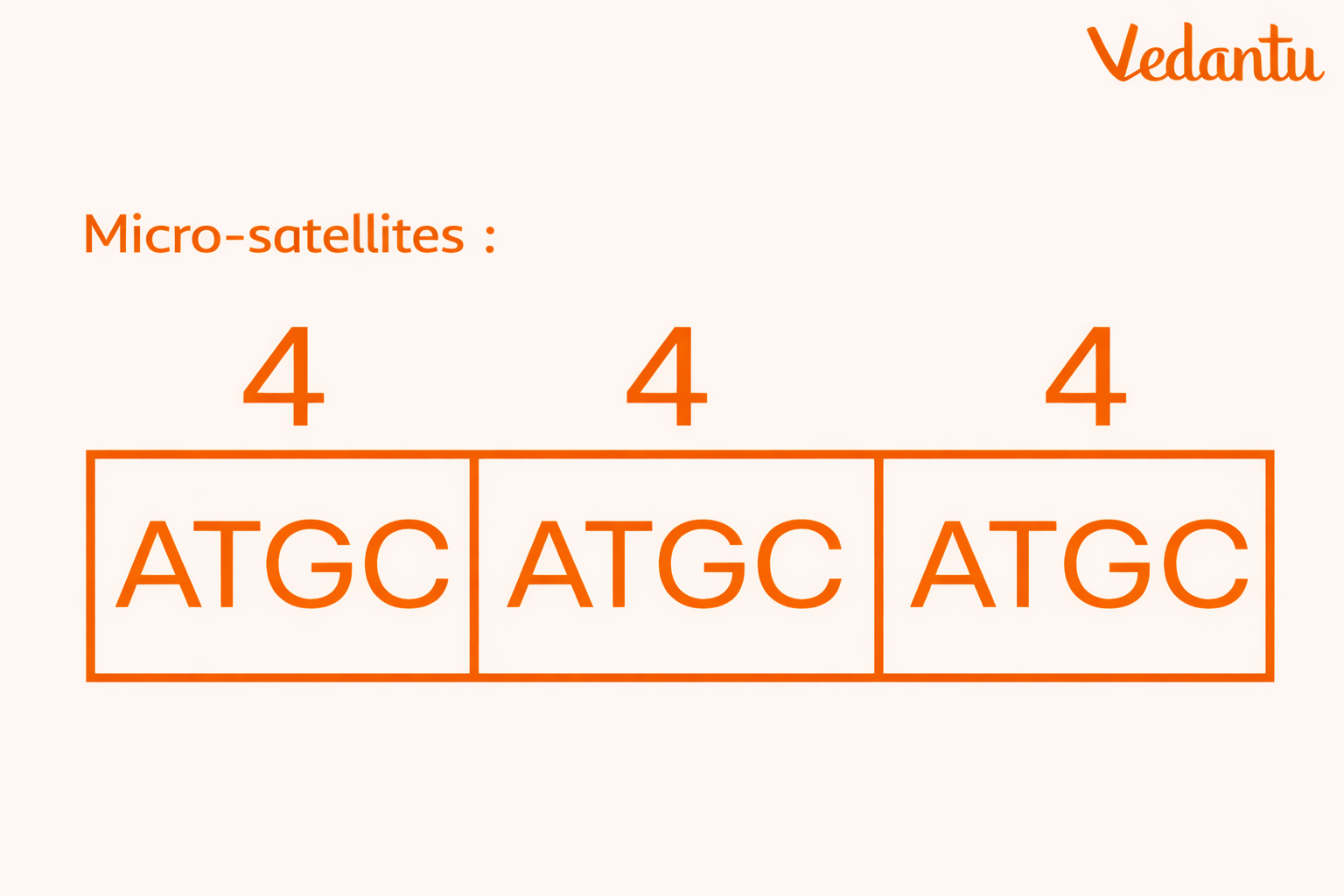

Microsatellites are short DNA sequences in which a unit of 1 to 6 base pairs is repeated many times.
They are also called SSR – Simple Sequence Repeats.
Key features of microsatellites
very short repeating units
present at multiple places in the genome
highly variable
useful in genetic analysis and DNA studies
Minisatellites
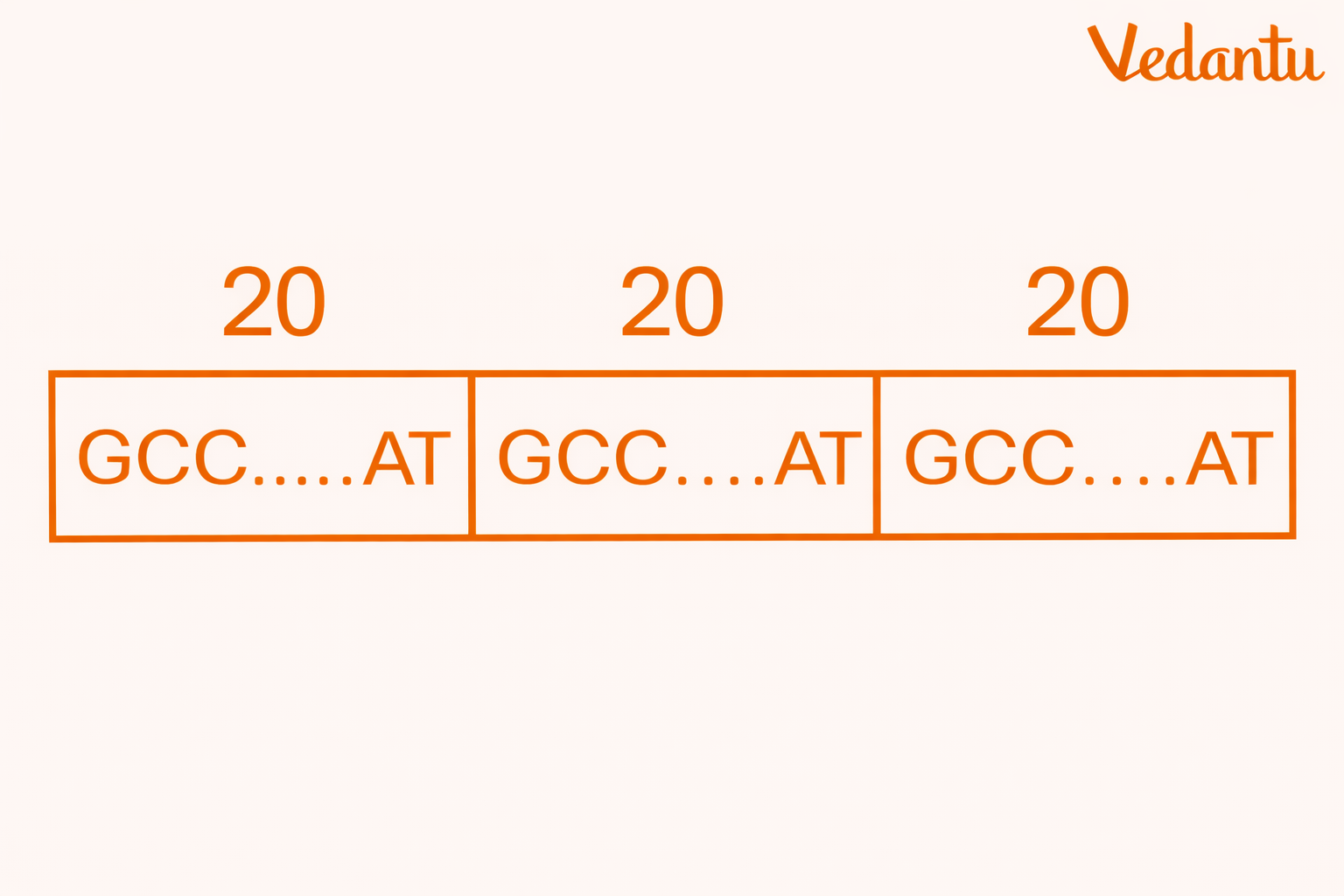
Minisatellites consist of repeated DNA units that are usually 11 to 60 base pairs long.
They are also called VNTRs – Variable Number of Tandem Repeats.
These sequences are extremely important in classical DNA fingerprinting because the number of repeat units varies from person to person, producing fragments of different lengths.
VNTR Full Form and Meaning
VNTR stands for Variable Number of Tandem Repeats.
In VNTR regions:
A short DNA sequence is arranged one after another in tandem
The number of copies varies between individuals
Even homologous chromosomes of the same individual may differ in repeat number
This variation produces different DNA fragment lengths
Important features of VNTR
Feature | Description |
Type | Minisatellite DNA |
Arrangement | Tandem repeats |
Variation | Copy number differs among individuals |
Size range | About 0.1 kb to 20 kb |
Importance | The main basis of DNA fingerprinting |
The high degree of polymorphism in VNTR regions makes them highly useful for identification.
RFLP in DNA Fingerprinting
RFLP stands for Restriction Fragment Length Polymorphism.
This is a variation in the length of DNA fragments produced when DNA from different individuals is cut with the same restriction enzyme.
.png)
Why Fragment Lengths Differ?
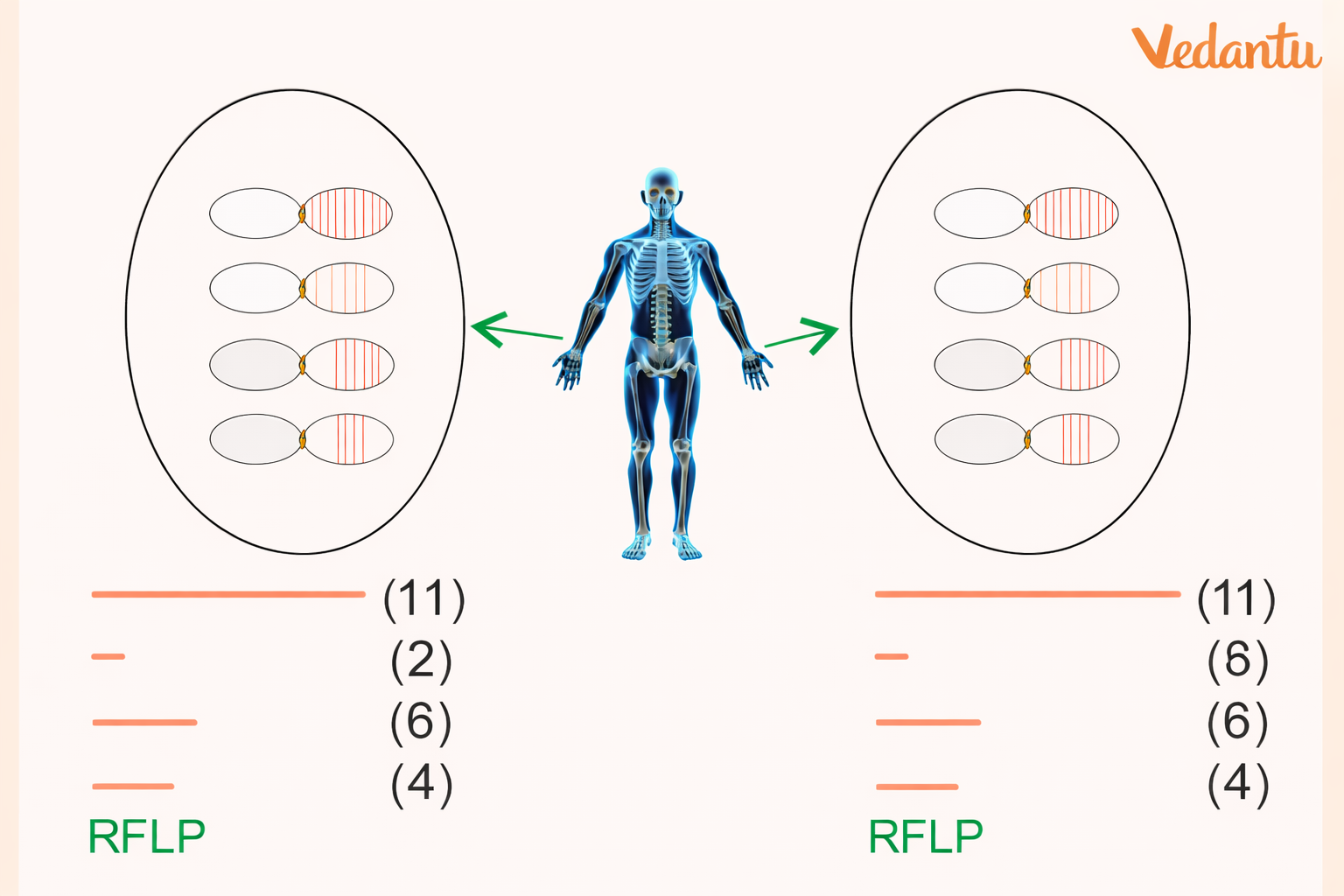
The number of VNTR repeats varies among individuals. So when DNA is cut using the same restriction enzyme, the resulting fragments are of different sizes.
These differences in fragment length create a distinctive banding pattern after hybridisation and autoradiography, and this pattern becomes the person’s DNA fingerprint.
RFLP Forms the Basis of DNA Fingerprinting
If two individuals have different VNTR repeat numbers, their DNA fragments will show:
different lengths
different band positions
different DNA fingerprint patterns
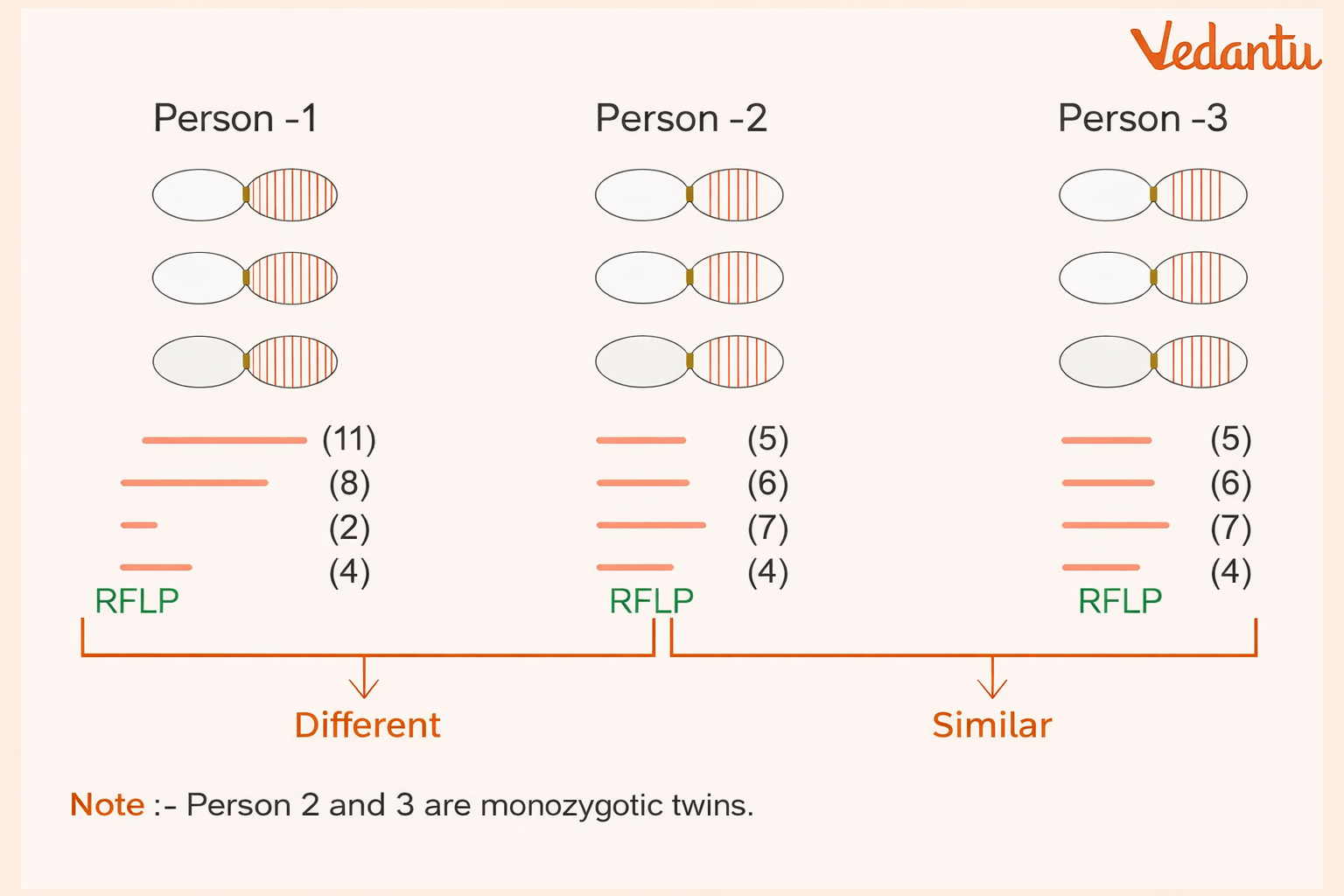
Cases where RFLP patterns can be the same
Monozygotic (identical) twins
Different tissues or cells of the same person
This is because identical twins originate from the same zygote and therefore have the same DNA pattern, and all tissues of one person contain the same DNA.
DNA Fingerprinting Technique: Steps of DNA Fingerprinting
DNA fingerprinting involves a sequence of carefully performed steps. Each step is important for obtaining an accurate and reliable DNA profile.
1. DNA Extraction
The first step is to isolate DNA from the biological sample. DNA may be obtained from:
blood
saliva
semen
hair root
skin cells
bone
tissues
The cells are broken open by cell lysis, and DNA is separated from the rest of the cellular material.
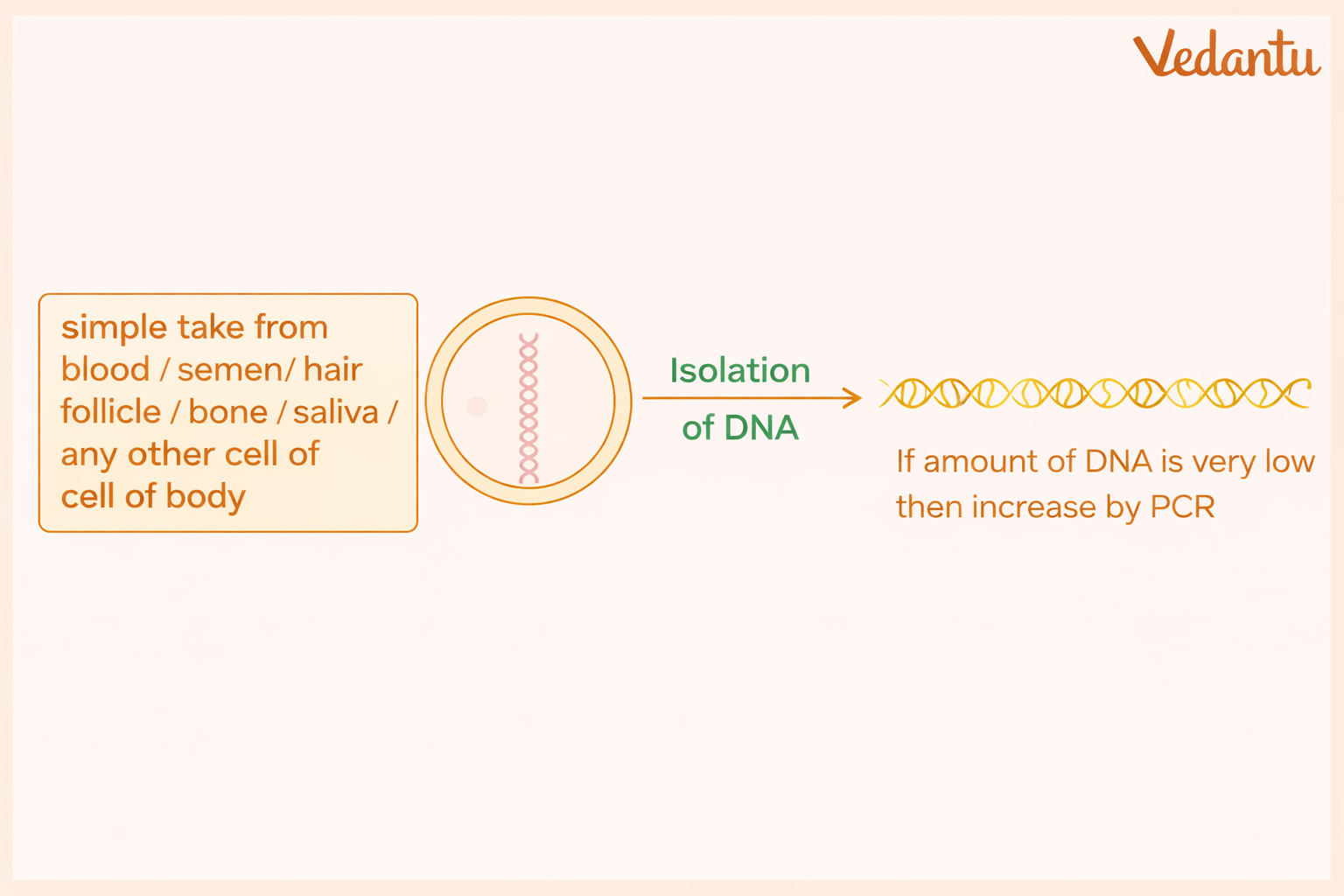
If the amount of DNA is very small, it may be amplified using PCR (Polymerase Chain Reaction) to generate enough DNA for analysis.
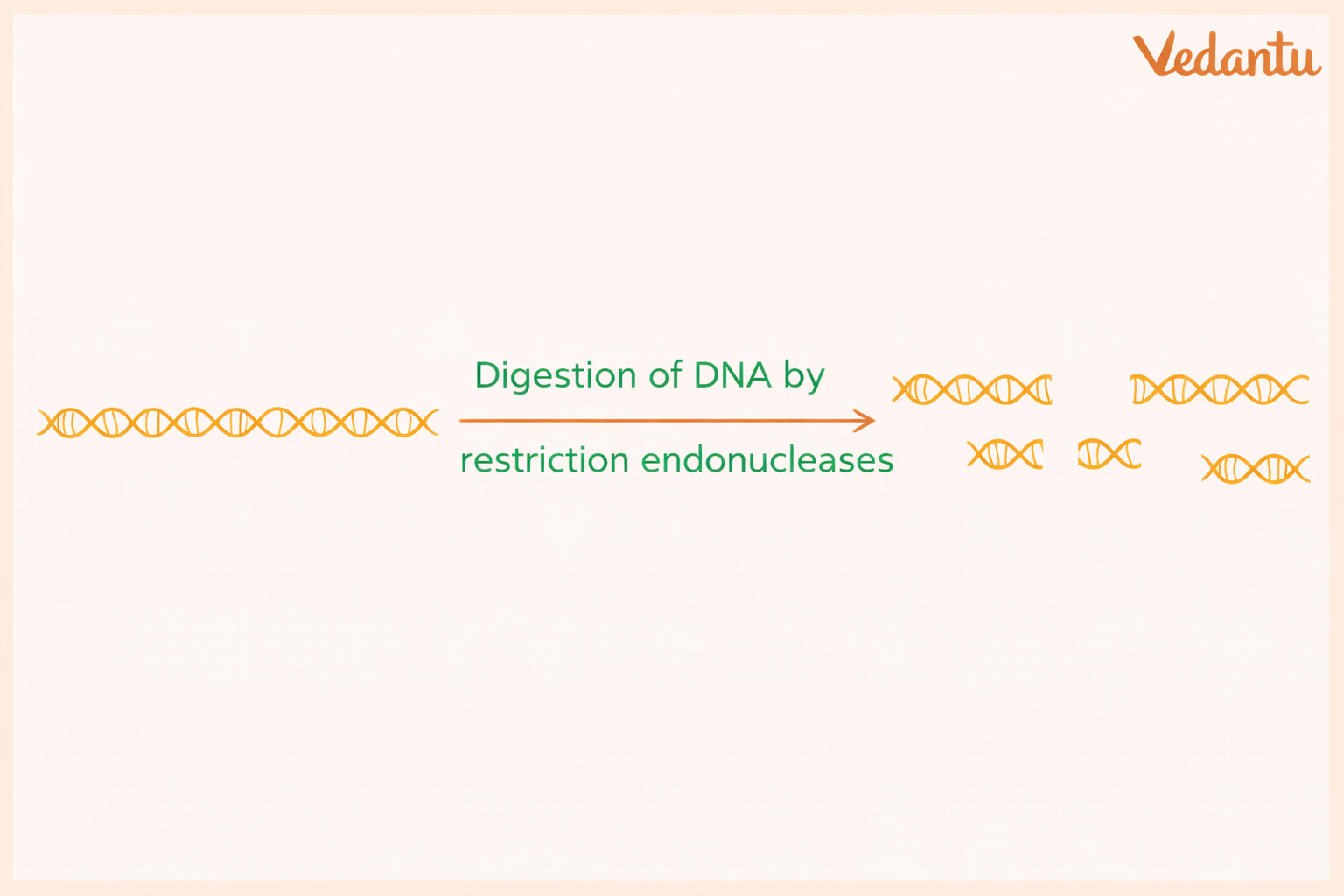
2. Restriction Enzyme Digestion
In this step, the extracted DNA is cut into fragments using a specific restriction enzyme.
The content mentions Hae III, an enzyme obtained from Haemophilus aegyptius, which recognises a specific DNA sequence, such as GGCC and cuts DNA at that site.
Because the number of repeats differs from person to person, the lengths of the cut fragments also differ. These fragments are then prepared for separation in a gel.
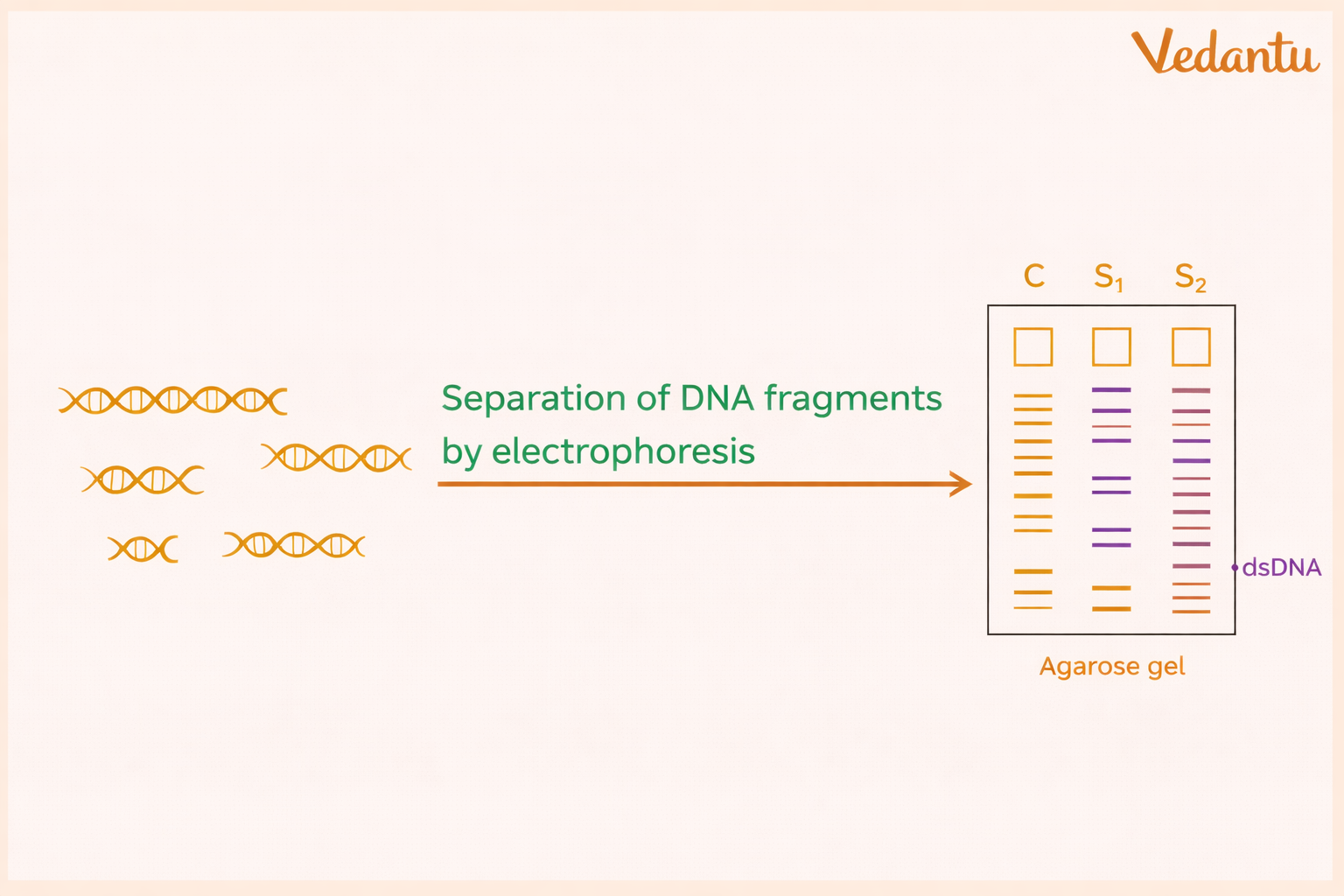
3. Gel Electrophoresis
The DNA fragments are separated using gel electrophoresis.
DNA carries a negative charge because of its phosphate backbone, so when an electric current is passed through the gel:
DNA fragments move toward the positive electrode
smaller fragments move faster
Larger fragments move more slowly
As a result, the DNA fragments become separated according to their size. Agarose gel is commonly used for this purpose.
4. Southern Blotting or Southern Transfer
After separation, the DNA fragments in the gel are transferred onto a nylon membrane. This technique is called Southern blotting, named after Edward Southern.
Before transfer, the DNA is denatured to form single strands. The separated DNA fragments are then blotted onto the membrane and fixed in position. This step stabilises the DNA, making it more suitable for hybridisation.
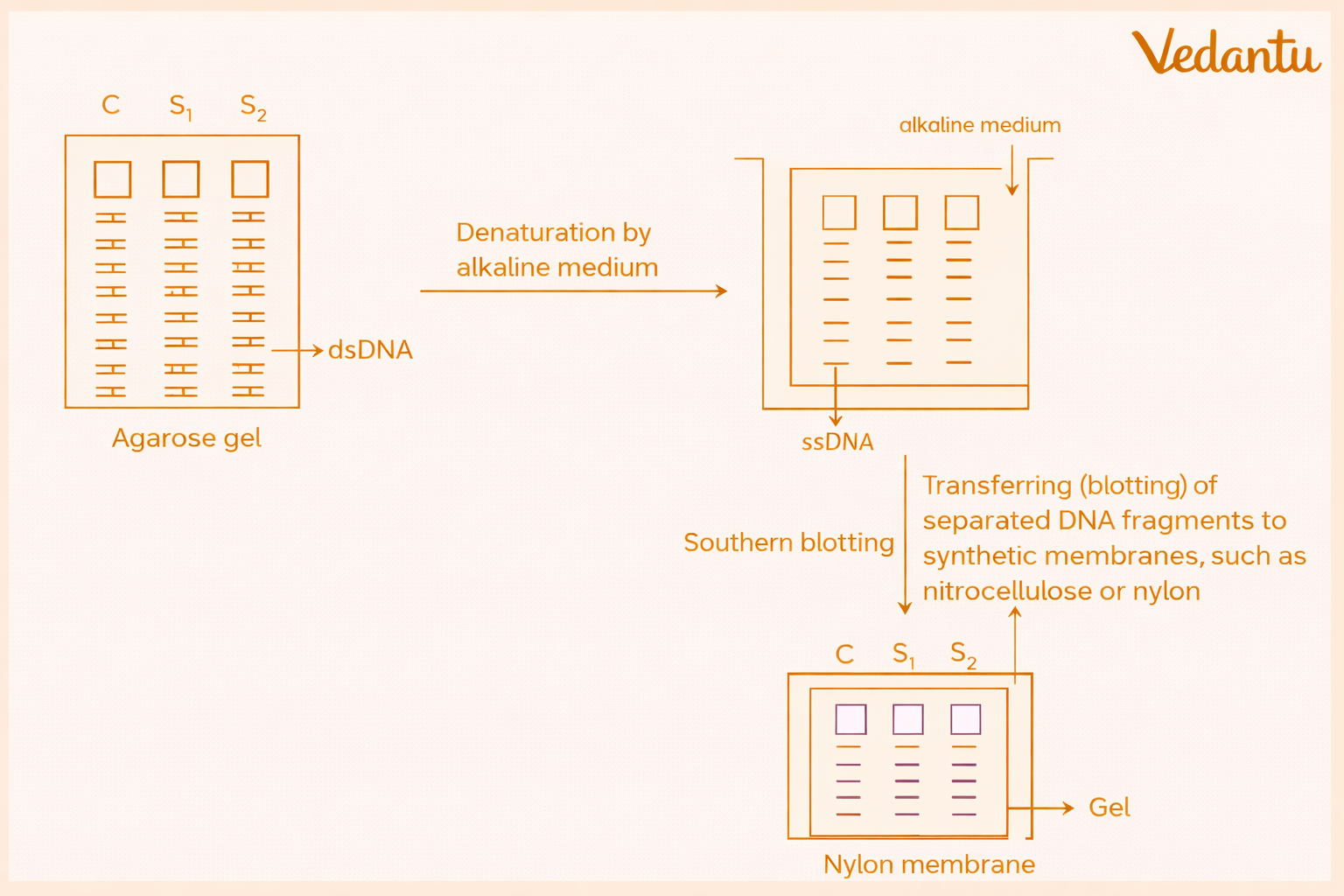
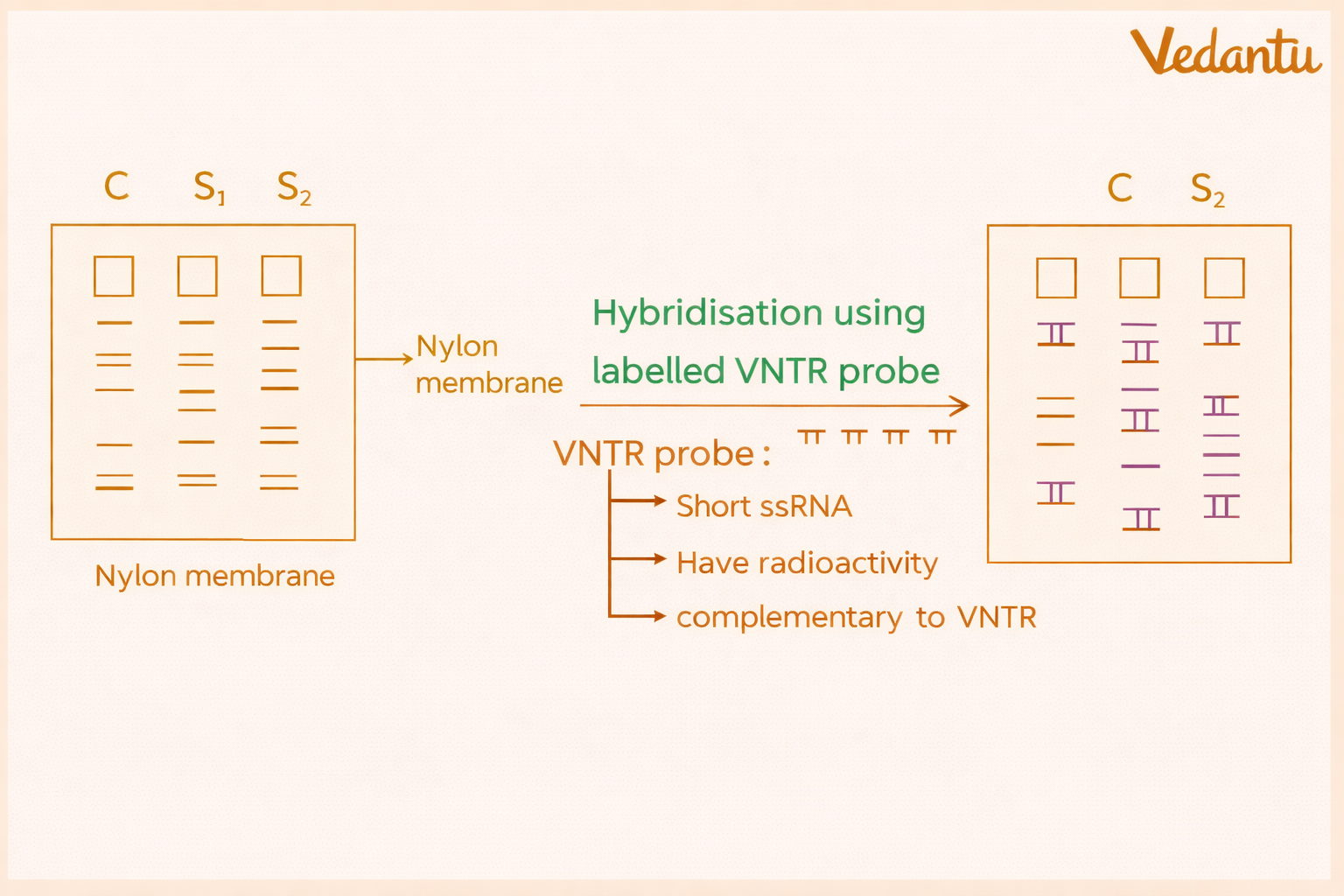
5. Hybridisation with Labelled Probe
In this step, a radioactively labelled DNA probe is used to detect specific DNA sequences, especially VNTR regions.
The probe is single-stranded and complementary to the target DNA sequence. It binds to matching DNA fragments on the membrane.
The content mentions probes labelled with radioactive phosphorus-32. These probes specifically bind to the variable repetitive DNA regions and help identify the positions of interest.
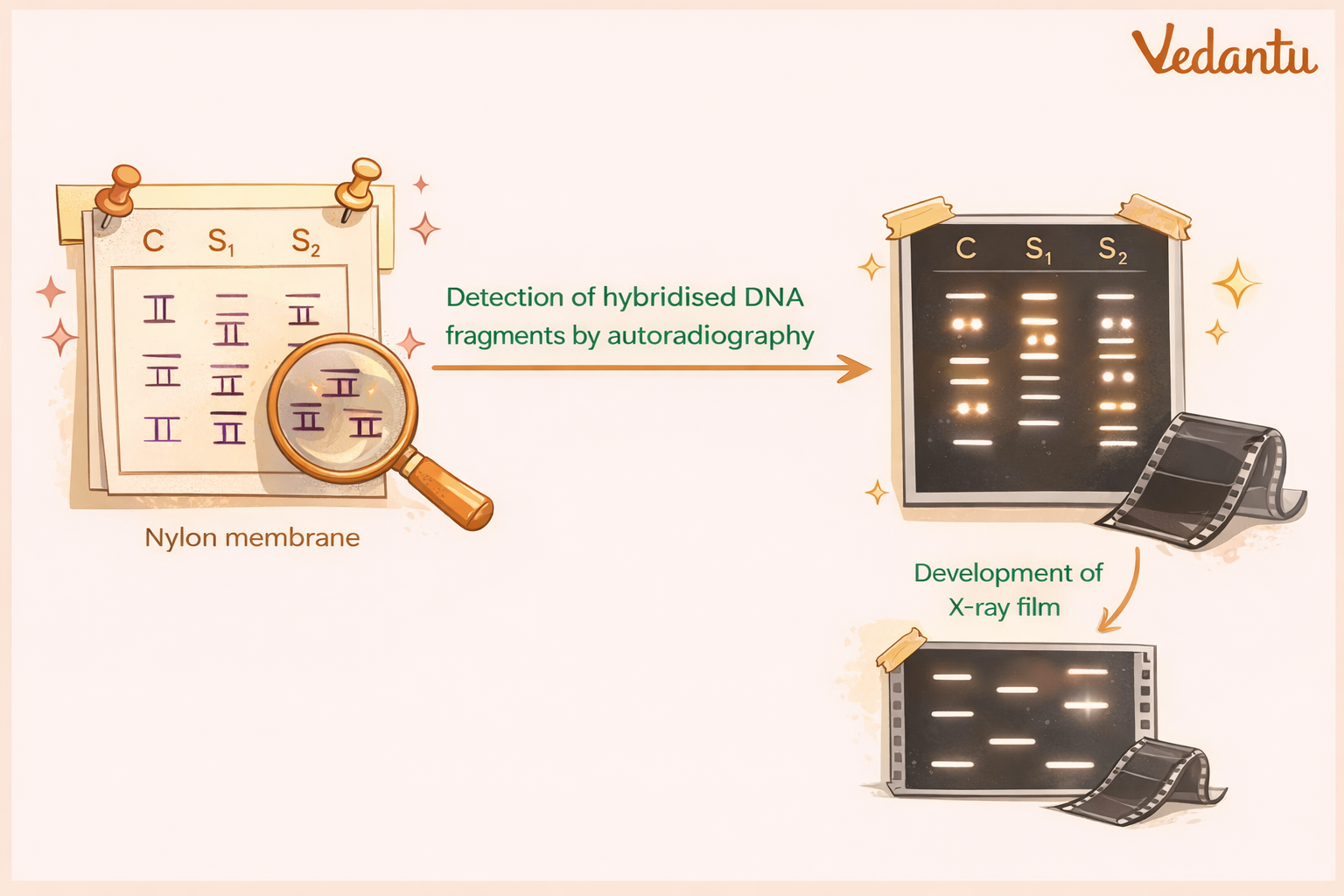
6. Autoradiography
The membrane containing the probe-bound DNA fragments is placed against an X-ray film.
The radioactive labels expose the film, producing dark bands at specific positions. These bands form a characteristic pattern called the DNA fingerprint.
This banding pattern:
is unique for each individual
Varies greatly among unrelated individuals
Is the same in all tissues of one person
Is the same in identical twins
DNA Fingerprinting Process at a Glance
Step | Process | Purpose |
1 | DNA extraction | Isolate DNA from cells |
2 | Restriction digestion | Cut the DNA into fragments |
3 | Gel electrophoresis | Separate fragments by size |
4 | Southern blotting | Transfer DNA to the membrane |
5 | Hybridisation | Bind probe to target sequence |
6 | Autoradiography | Visualise band pattern |
This sequence produces the final DNA profile used for identification.
Why is DNA Fingerprinting Highly Reliable?
DNA fingerprinting is considered highly reliable because the chance of two unrelated individuals having the same fingerprint is extremely low.
Its reliability comes from:
high polymorphism in VNTR regions
inheritance of DNA patterns
stability of DNA in different tissues
precise laboratory methods
This makes DNA fingerprinting a powerful identification tool in biology, medicine, and forensic science.
Applications of DNA Fingerprinting
DNA fingerprinting has a wide range of practical applications.
1. Paternity Testing
This is one of the most common uses of DNA fingerprinting. The DNA fingerprint of a child is compared with that of the mother and the alleged father.
Since a child inherits:
50% of the DNA from the mother
50% of the DNA from the father
Matching patterns help determine biological parentage. DNA fingerprinting is therefore widely used in paternity disputes and family relationship testing.
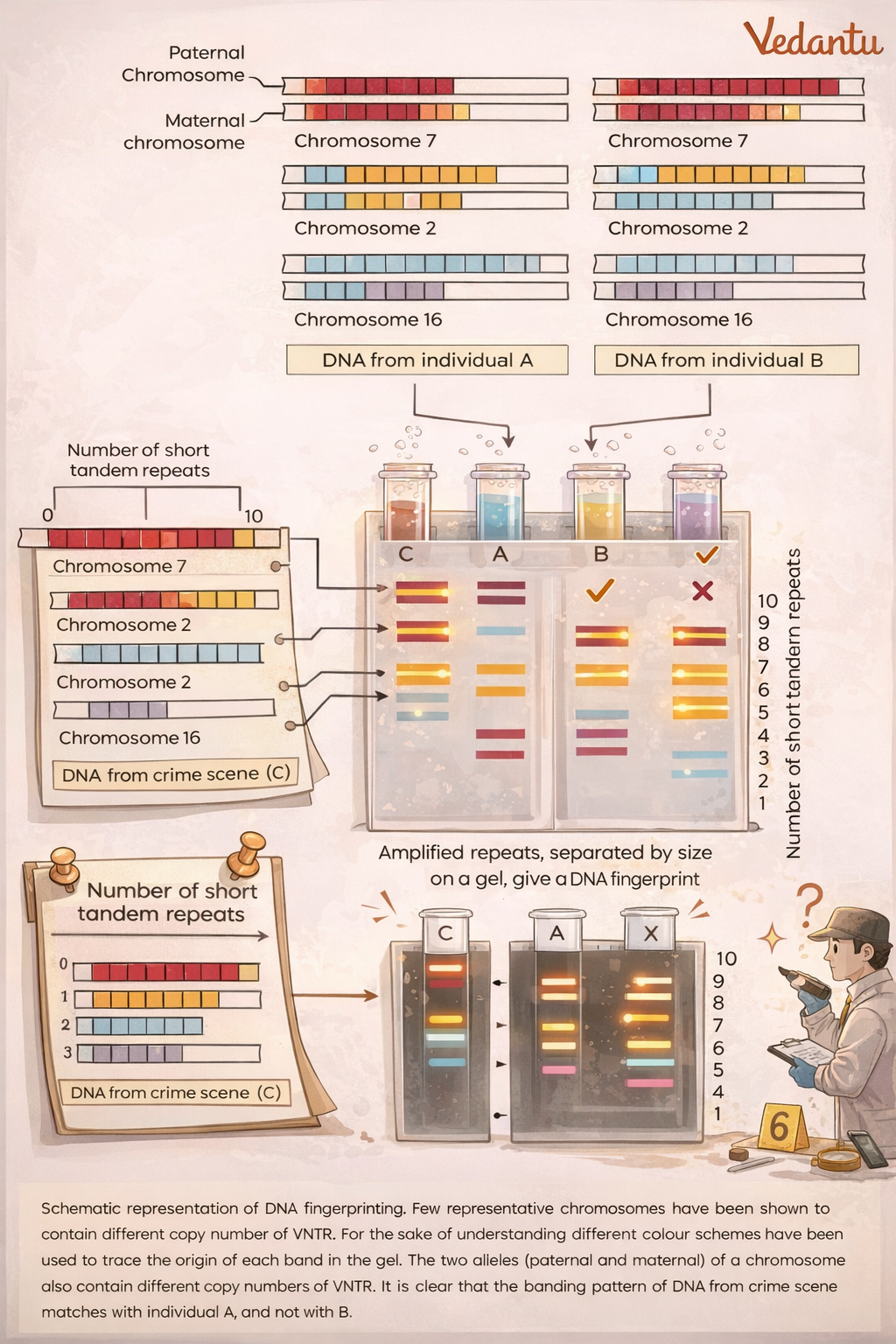
2. Criminal Identification
DNA fingerprinting plays a major role in forensic science.
Biological samples collected from a crime scene, such as:
blood
semen
hair roots
saliva
skin tissue
It can be compared with the suspects' DNA.
This is especially useful in serious cases such as:
murder
rape
assault
missing persons cases
By matching DNA patterns, investigators can connect a suspect to the crime scene with a high degree of accuracy.
3. Identification of Dead Bodies
DNA fingerprinting is extremely useful in identifying bodies in cases of:
natural disasters
air crashes
fires
accidents
war casualties
Even when the body is damaged or difficult to identify visually, DNA can help establish identity by comparing samples with those of family members.
4. Immigration and Family Disputes
DNA fingerprinting is also used in legal and civil cases where biological relationships need to be proven. It can help in:
immigration disputes
inheritance cases
child custody matters
family reunification claims
5. Population and Genetic Diversity Studies
DNA fingerprinting helps scientists study:
genetic variation within populations
relatedness among individuals
biodiversity
population structure
evolutionary relationships
By comparing DNA patterns among groups, researchers can understand how genetically similar or different populations are.
Importance of DNA Fingerprinting in India
In India, CDFD – Centre for DNA Fingerprinting and Diagnostics, Hyderabad, is an important institution associated with DNA fingerprinting research and applications.
It has played a significant role in:
DNA-based diagnostics
forensic applications
genetic studies
research and training
DNA Fingerprinting vs DNA Sequencing
Students often confuse DNA fingerprinting with DNA sequencing, so this distinction is useful.
Basis | DNA Fingerprinting | DNA Sequencing |
Purpose | Identify individuals | Determine exact nucleotide order |
DNA analysed | Selected variable regions | Entire sequence or target sequence |
Speed | Faster for identification | More detailed but time-consuming |
Cost | Less expensive for routine identity work | Usually more expensive |
Main use | Forensics, paternity, identification | Genomics, mutation analysis, research |
DNA fingerprinting does not require complete DNA sequencing. It only studies selected highly variable repetitive regions.
Key Terms in DNA Fingerprinting
Term | Meaning |
DNA profiling | Another name for DNA fingerprinting |
Polymorphism | Variation in the DNA sequence among individuals |
Repetitive DNA | DNA sequence repeated many times |
Satellite DNA | Highly repetitive non-coding DNA |
Microsatellite | 1–6 bp repeats, also called SSR |
Minisatellite | 11–60 bp repeats, also called VNTR |
VNTR | Variable Number of Tandem Repeats |
RFLP | Restriction Fragment Length Polymorphism |
Probe | Labelled DNA sequence used to detect target DNA |
Southern blotting | Transfer of DNA from gel to membrane |
Autoradiography | X-ray film detection of radioactive bands |
DNA Fingerprinting Flowchart
Sample collection → DNA extraction → Restriction digestion → Gel electrophoresis → Southern blotting → Probe hybridisation → Autoradiography → DNA band pattern analysis
This is the complete classical workflow of DNA fingerprinting.
Exam-Focused Points to Remember
DNA fingerprinting is based on polymorphism in repetitive DNA sequences
It was developed by Sir Alec Jeffreys in 1984
The most important repetitive sequences used are VNTRs
VNTR belongs to minisatellite DNA
RFLP is the variation in fragment length after restriction digestion
DNA from all tissues of one individual shows the same fingerprint
Identical twins have the same DNA fingerprint
DNA fingerprinting is used in forensics, paternity testing, and population studies
In India, an important centre is CDFD, Hyderabad
Conclusion
DNA fingerprinting is one of the most significant tools in modern biology and forensic science. It is based on variations in repetitive DNA regions, especially VNTRs, and helps create a unique genetic profile for every individual except identical twins. From crime investigation and paternity testing to population genetics and body identification, DNA fingerprinting has transformed the way biological identity is established. For students, this topic is important because it integrates genetics, molecular biology, biotechnology, and real-world applications in a highly exam-relevant way.
Also Read – Related Topics for DNA Fingerprinting
FAQs on DNA Fingerprinting Explained: Principle, Steps, Diagrams, and Applications
1. What is DNA fingerprinting in simple words?
DNA fingerprinting is a technique used to identify a person by analysing unique patterns in their DNA.
2. Who discovered DNA fingerprinting?
DNA fingerprinting was discovered by Sir Alec Jeffreys in 1984.
3. What is the principle behind DNA fingerprinting?
It is based on polymorphism in repetitive DNA regions, especially VNTRs, which vary from one individual to another.
4. What is a VNTR in DNA fingerprinting?
VNTR stands for Variable Number of Tandem Repeats. These are minisatellite DNA regions in which the number of repeated sequences differs among individuals.
5. What is RFLP?
RFLP stands for Restriction Fragment Length Polymorphism. It refers to differences in the lengths of DNA fragments produced after restriction enzyme digestion.
6. Why is DNA fingerprinting important in forensic science?
It helps identify criminals by matching biological samples from a crime scene to suspects' DNA.
7. Can DNA fingerprinting be used for paternity testing?
Yes. It compares the DNA of the child, mother, and possible father to determine biological parentage.
8. Are DNA fingerprints the same in all tissues of a person?
Yes. Blood, hair roots, skin, bone, saliva, and other tissues from the same individual share the same DNA fingerprint.
9. Do identical twins have the same DNA fingerprint?
Yes. Monozygotic twins have the same DNA fingerprint because they originate from the same zygote.
10. Which organisation in India is associated with DNA fingerprinting?
CDFD, Hyderabad, is an important Indian institution for DNA fingerprinting and diagnostics.
11. What are the 6 steps of DNA fingerprinting?
The steps of DNA fingerprinting are usually explained in six main stages. In the DNA Fingerprinting Class 12, these steps are important for understanding the full DNA fingerprinting process and its principles.
DNA Extraction
DNA is first isolated from the sample. This sample may come from blood, hair root, saliva, semen, or other body cells. If the amount of DNA is very small, it can be amplified using PCR.Restriction Digestion
The isolated DNA is cut into smaller pieces using restriction enzymes. These enzymes cut DNA at specific sites and produce fragments of different lengths.Gel Electrophoresis
The DNA fragments are separated on an agarose gel according to size. Smaller fragments move faster, while larger fragments move more slowly.Southern Blotting
The separated DNA fragments are transferred from the gel to a nylon or nitrocellulose membrane.Hybridisation
The membrane is treated with labelled probes that bind to specific repetitive DNA sequences such as VNTR regions.Autoradiography
The final band pattern is seen on X-ray film. This band pattern forms the DNA fingerprinting diagram or profile of the individual.
These six stages together make up the DNA fingerprinting technique.
12. When was DNA discovered in 1984?
In the context of DNA fingerprinting, the important date is 10 September 1984, when Sir Alec Jeffreys discovered it. On that day, Alec Jeffreys observed a unique banding pattern in DNA that led to the discovery of DNA fingerprinting.
13. What are three ways DNA fingerprinting is used?
The main applications of DNA fingerprinting include:
Criminal investigations
It is used to match DNA from a crime scene with a suspect.Paternity testing
It helps find biological relationships between parents and children.Personal identification
It is used to identify unknown persons, missing people, or bodies in disaster cases.
14. Who started DNA fingerprinting in India?
Lalji Singh is known as the person who started DNA fingerprinting in India. He is widely regarded as the Father of Indian DNA Fingerprinting for his major contribution to bringing and developing this technology in the country.
15. How old is the oldest known DNA?
The oldest known DNA successfully studied is about 2.4 million years old. Scientists have recovered and sequenced this ancient DNA from very old biological material.
This fact is not directly part of the DNA fingerprinting steps, but it shows how powerful DNA analysis has become in modern science.
16. What is DNA fingerprinting called nowadays?
Nowadays, DNA fingerprinting is more commonly called DNA profiling. It is also sometimes called genetic fingerprinting or DNA typing.
Even though the name has changed in many modern uses, the meaning remains the same. It is a DNA fingerprinting technique used to identify individuals by studying unique DNA patterns.
17. Why is 98% of our DNA called junk DNA?
A large part of human DNA was once called junk DNA because it does not encode proteins. Scientists once believed that these non-coding regions had no major function.























